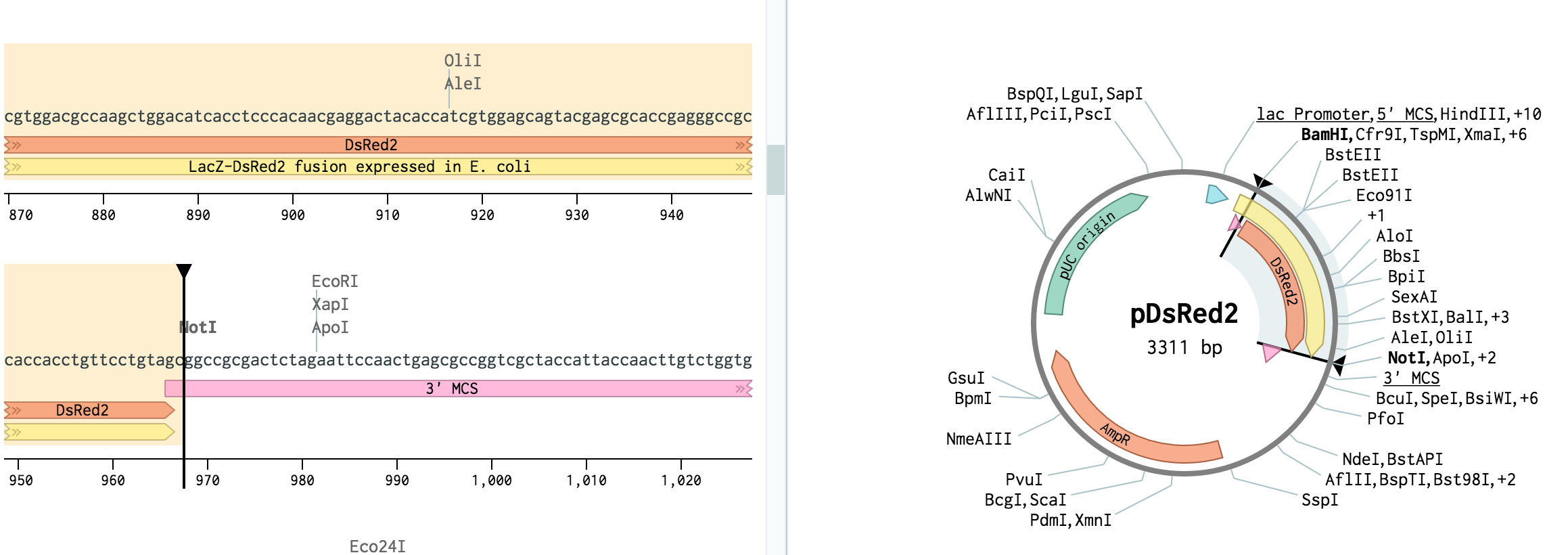
Tip: Methylation patterns also differ between different eukaryotes (see bullets above), affecting the choice of restriction enzyme for construction genomic DNA libraries. The frequency of cutting in a random DNA sequence for a given restriction enzyme is once per every 4n, where n is the number of bases in the restriction enzymes. Tip: Methylation patterns differ between bacteria and eukaryotes, so restriction patterns of cloned and uncloned DNA may differ. Plant DNA is highly methylated, so for successful mapping in plants, choose enzymes that either do not contain a CpG dinucleotide in their recognition site (e.g., DraI or SspI) or that can cleave methylated CpG dinucleotides (e.g., BamHI, KpnI, or TaqI).Rare-cutter enzymes therefore cleave more frequently in these species. Drosophila, Caenorhabditis, and some other species do not possess methylated DNA, and have a higher proportion of CpG dinucleotides than mammalian species.Therefore many enzymes with CpG in their recognition site, such as EagI, NotI, and SalI, cleave mammalian DNA only rarely. The CpG dinucleotide occurs about 5 times less frequently in mammalian DNA than would be expected by chance, and most restriction enzymes with a CpG dinucleotide in their recognition site do not cleave if the cytosine is methylated. 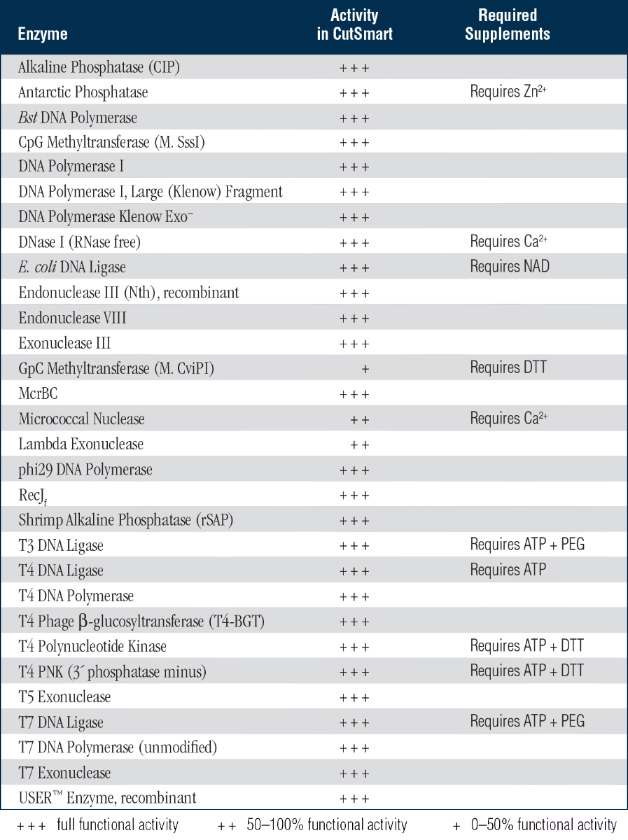 
And a PCR product, made by Taq (primers not phosphorylated). In addition, methylation patterns differ in different species, also affecting the choice of restriction enzyme. Blunt ligation with PCR: is kinase needed - posted in Molecular Cloning: Hi, sorry for the question, but Im somehow confused here. Therefore the choice of restriction enzyme is affected by its sensitivity to methylation. Not all restriction enzymes can cleave their recognition site when it is methylated. Many organisms have enzymes called methylases that methylate DNA at specific sequences.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
